
However, in the past 30 years, significant advances in isotope labeling, NMR pulse sequence development and software methods paved the path for multidimensional NMR studies of biomacromolecules. The funders had no role in study design, data collection and analysis, decision to publish, or preparation of the manuscript.Ĭompeting interests: The authors have declared that no competing interests exist.Įarly NMR studies of biomacromolecules were performed on relatively simple low molecular weight model systems using one-dimensional (1D) methods. This is an open access article distributed under the terms of the Creative Commons Attribution License, which permits unrestricted use, distribution, and reproduction in any medium, provided the original author and source are creditedĭata Availability: All relevant data are within the paper and its Supporting Information files.įunding: This work was supported by National Institutes of Health (NIH) National Research Service Award (NRSA) F32 DK097890 (T.S.H.), NIH Pathway to Independence Award K99 DK103116 (T.S.H.) NIH R01 grant GM114420 (D.J.K.) NIH R01 grant DK101871 (D.J.K.) a New Investigator Award 1KN-09 from the James and Esther King Biomedical Research Program, Florida Department of Health (D.J.K.) and the State of Florida to Scripps Florida via institutional start-up funds (D.J.K.). Received: FebruAccepted: JPublished: August 4, 2015Ĭopyright: © 2015 Hughes et al. Driscoll, The Francis Crick Institute, UNITED KINGDOM The decon1d program is freely available as a downloadable Python script at the project website ( ).Ĭitation: Hughes TS, Wilson HD, de Vera IMS, Kojetin DJ (2015) Deconvolution of Complex 1D NMR Spectra Using Objective Model Selection. In contrast, the BIC method used by decond1d provides a quantitative method for model comparison that penalizes for model complexity helping to prevent over-fitting of the data and allows identification of the most parsimonious model. In current methods, determination of the deconvolution model best supported by the data is done manually through comparison of residual error values, which can be time consuming and requires model selection by the user. The method also allows for fitting of intermediate exchange spectra, which is not supported by current software in the absence of a specific kinetic model.
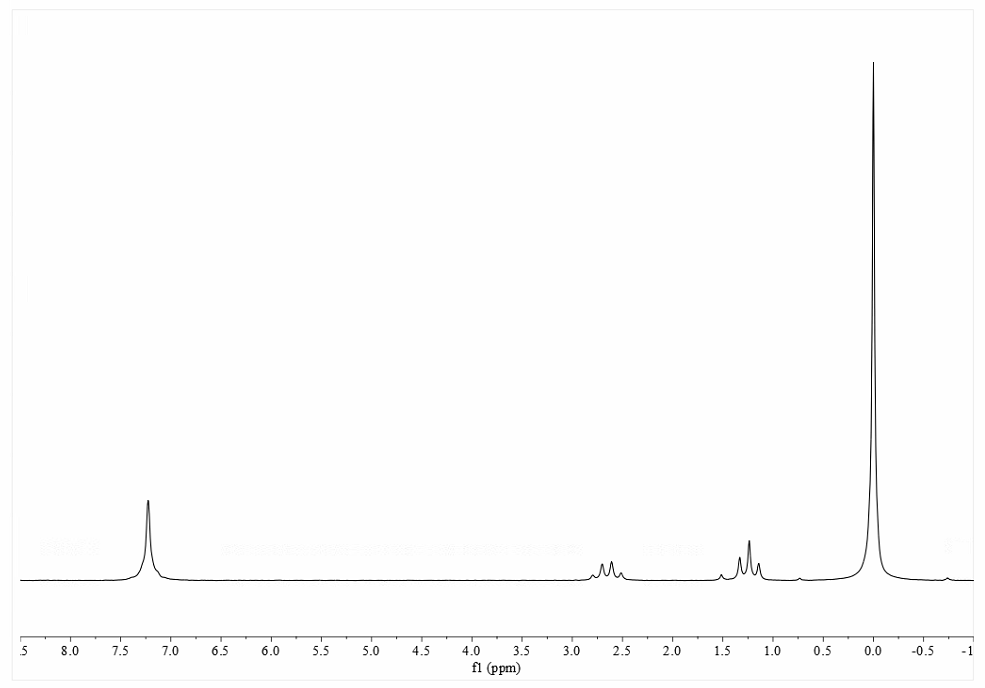
We have developed a Python-based deconvolution program, decon1d, which uses Bayesian information criteria (BIC) to objectively determine which model (number of peaks) would most likely produce the experimentally obtained data. One-dimensional (1D) 19F NMR spectra of proteins can be broad, irregular and complex, due to exchange of probe nuclei between distinct electrostatic environments and therefore cannot be deconvoluted and analyzed in an objective way using currently available software.
Fluorine ( 19F) NMR has emerged as a useful tool for characterization of slow dynamics in 19F-labeled proteins.
